Dr. Kannan: New UGA study maps the structural diversity of sugar building enzymes using AI
Tuesday, October 5, 2021 – 10:22am
By:Alan Flurry
Sugars in our bodies, and in nearly all living organisms, are synthesized and built by a large family of proteins called Glycosyltransferases (GTs) that adopt unique three-dimensional structures and folds to perform a diverse array of cellular functions. Understanding the structure and fold of these proteins is an important first step towards characterizing their functions, which is critical for developing effective glycovaccines and for improving crop yields and sustainable biofuels through the synthetic design of glycoproteins with desirable functional properties.
However, despite significant efforts in the structural characterization of GTs, mapping the full functional and fold landscape remains a challenge because of the large and diverse nature of these proteins and the cost and time associated with their structural characterization.
To address this challenge, an inter-disciplinary team of UGA researchers have leveraged recent advances in deep learning to predict and classify GT folds from primary sequences with high accuracy. Deep learning is a branch of artificial intelligence (AI) that uses interconnected artificial neural networks to automatically find patterns in large datasets.
The methods are designed to mimic the learning process of the human brain and are widely used in a range of applications from marketing to self-driving cars. However, their application in biology is only being realized, thanks to the massive amounts of biological data generated from gene sequencing studies.
“With over a half million GT sequences available, investigating the relationships connecting primary sequence, fold and function is a problem well poised for the application of deep learning methods” said Natarajan Kannan, professor of biochemistry and molecular biology and the Institute of Bioinformatics in the Franklin College of Arts and Sciences who led the research team.
Their work, published in the journal Nature Communications, reports the development of an “interpretable” deep learning model for predicting GT fold and function from primary sequences.
“One unique aspect of our model is that it is simple and interpretable, meaning that the neural network can be tracked by identifying the neurons that get activated during the learning process, which, in turn, helps in the biological interpretation of the prediction and classification process,” said Sheng Li, assistant professor of computer science and co-author on the study. “This is conceptually different from most existing deep learning models that operate as a “black box.”
“By predicting GTs that can adopt novel folds, this study provides a range of structural templates along with their crucial functional features for the design and synthesis of novel GTs for various applications,” said Kannan, who received a Maximizing Investigator Research Award, or MIRA, from the National Institute of General Medical Sciences in March 2021.
“This method promises to be a valuable tool for the glycobiology community and marks a significant milestone towards leveraging the full potential of GTs in biomedicine and other industries. It’s one great advantage to receiving the NIGMS award, which provides us with the flexibility to move in new and exciting directions,” he said.
The award and the new study help support interdisciplinary graduate training in the Institute of Bioinformatics and the department of computer science, as well build new synergies with campus-wide AI initiatives at UGA.
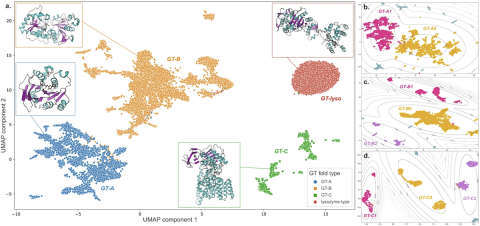
Image: Figure 2 from the study. Dots represent 2D UMAP projection of features for individual sequences.
Chronicles Tagging:
